SVM
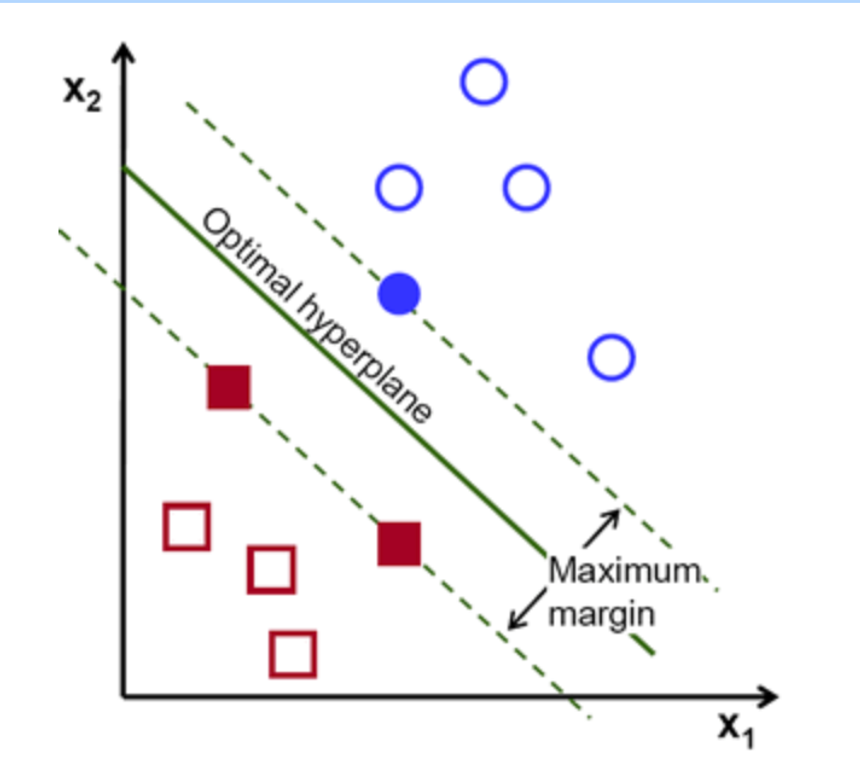
Definition of a hyperplane:
Canonical hyperplane: by convention, among all the possible, we choose , where represent the training image closest to the hyperplane
Distance between a point and hyperplane:
The margin M is twice the distance to the closest example:
Finally, maximizing M is minimizing with some constraints to correctly classify all training examples
subject to
this is a Lagrangian optimization problem
full script
Python
Copy
labels = np.array([1, -1, -1, -1])
trainingData = np.matrix([[501, 10], [255, 10], [501, 255], [10, 501]], dtype=np.float32)
# train
svm = cv.ml.SVM_create()
svm.setType(cv.ml.SVM_C_SVC)
svm.setKernel(cv.ml.SVM_LINEAR)
svm.setTermCriteria((cv.TERM_CRITERIA_MAX_ITER, 100, 1e-6))
svm.train(trainingData, cv.ml.ROW_SAMPLE, labels)
# Data for visual representation
width = 512
height = 512
image = np.zeros((height, width, 3), dtype=np.uint8)
green = (0,255,0)
blue = (255,0,0)
for i in range(height):
for j in range(width):
sampleMat = np.matrix([[j,i]], dtype=np.float32)
response = svm.predict(sampleMat)[1]
if response == 1:
image[i,j] = green
elif response == -1:
image[i,j] = blue
thickness = 2
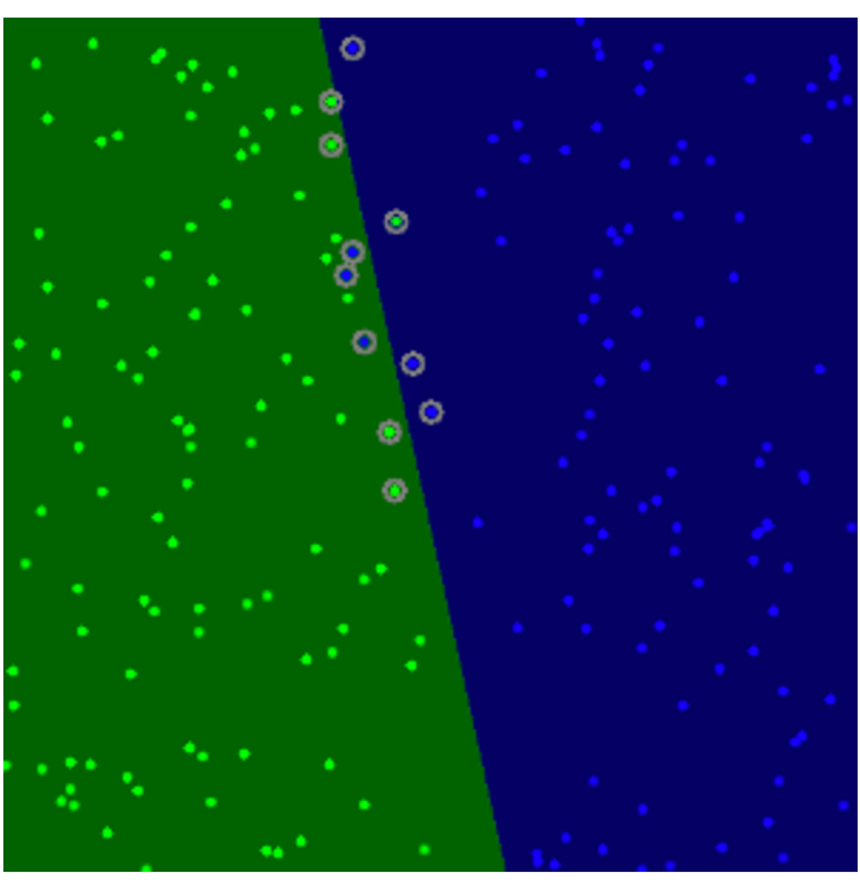
sv = svm.getUncompressedSupportVectors()
for i in range(sv.shape[0]):
cv.circle(image, (sv[i,0], sv[i,1]), 6, (128, 128, 128), thickness)
Non Linear SVM
The training data can be rarely separated using an hyperplane
Chance is more for a non-linear separable data in lower-dimensional space to become linear separable in higher-dimensional space.
In general, it is possible to map points in a d-dimensional space to some D-dimensional space to check the possibility of linear separability
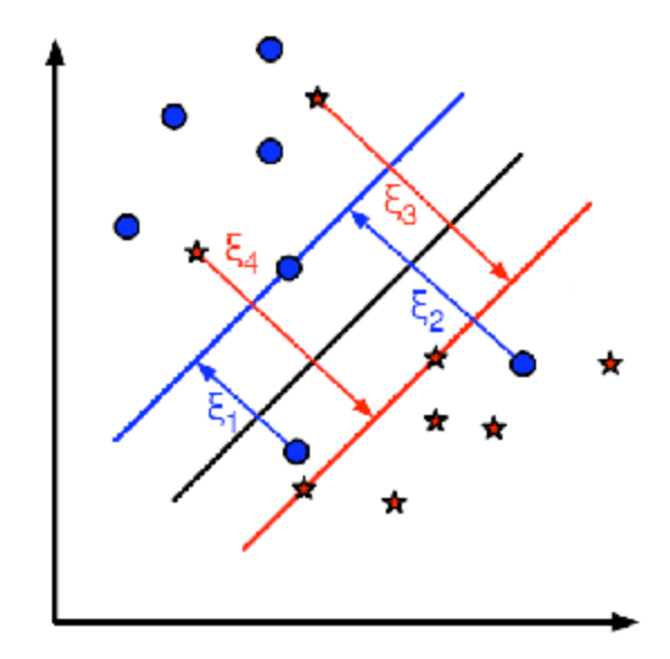
The new model has to include both the old requirement of finding the hyperplane that gives the biggest margin and the new one of generalizing the training data correctly by not allowing too many classification errors.
For example, one could think of minimizing the same quantity plus a constant times the number of misclassification errors in the training data
A better solution will take into account the distance of the misclassified samples to their correct decision regions
subject to
and
Choose C
Large values give solutions with less classification errors but smaller margin
Small values of C give solutions focusing more on the hyperplane, without much importance for classification errors, so wider margin
Implementation
Python
Copy
svm = cv.ml.SVM_create()
svm.setType(cv.ml.SVM_C_SVC)
svm.setC(0.1) # new
svm.setKernel(cv.ml.SVM_LINEAR)
svm.setTermCriteria((cv.TERM_CRITERIA_MAX_ITER, int(1e7), 1e-6))
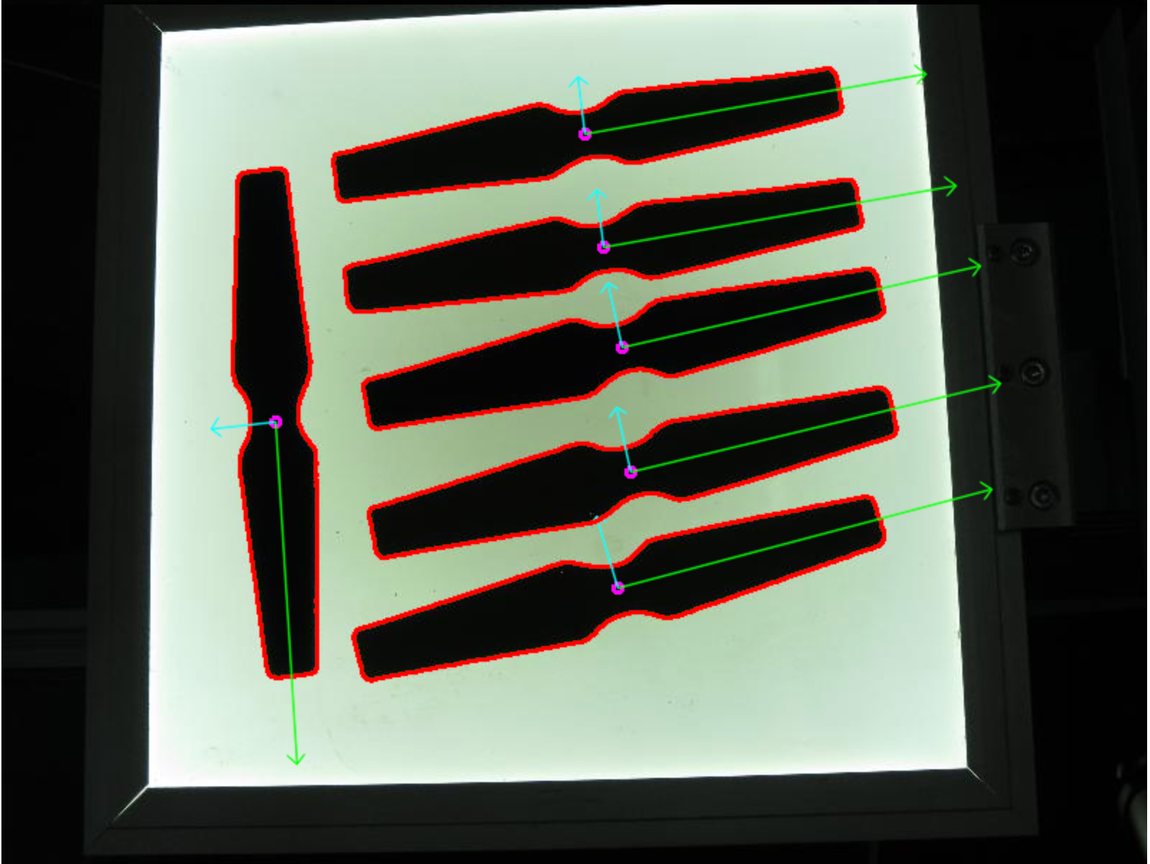
PCA
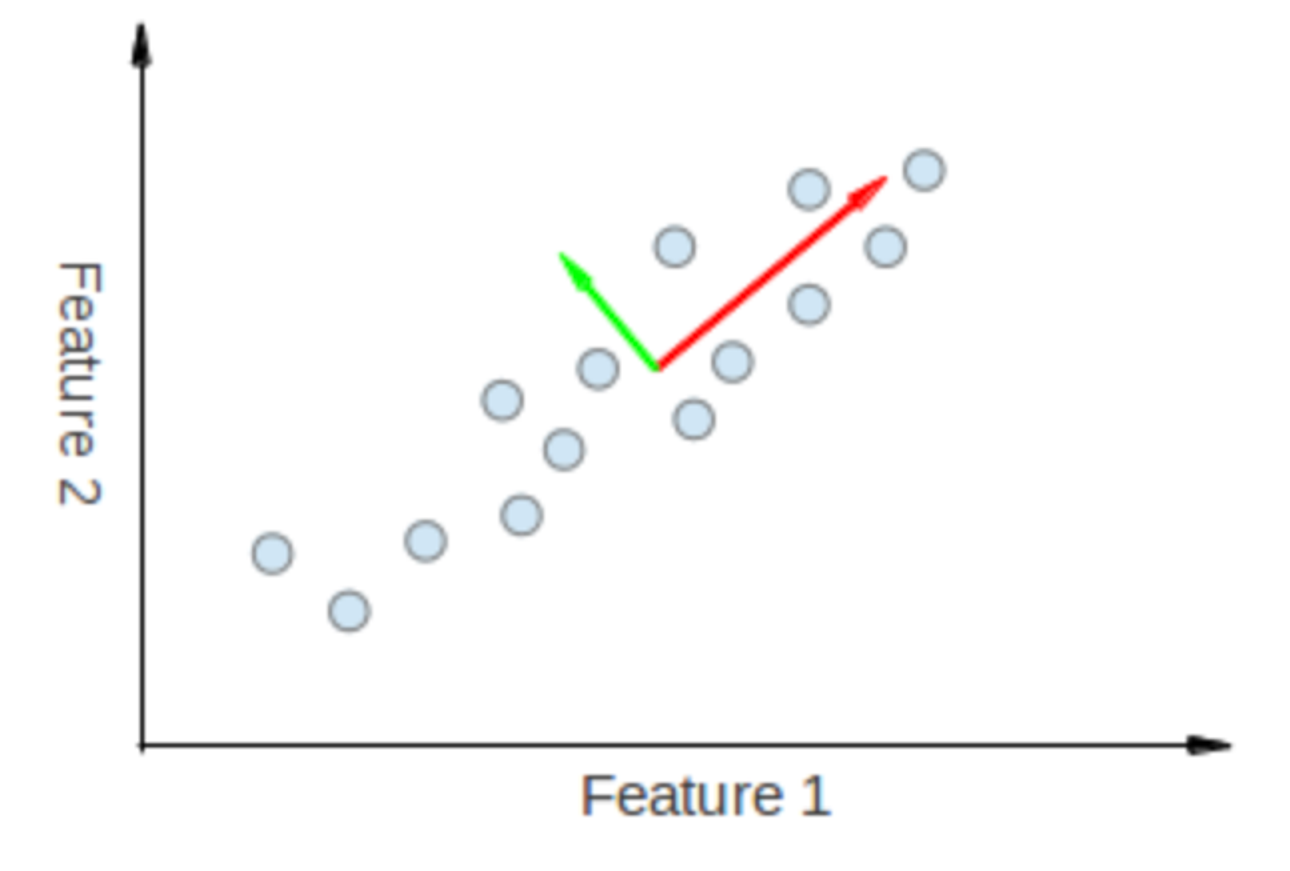
eigenvectors
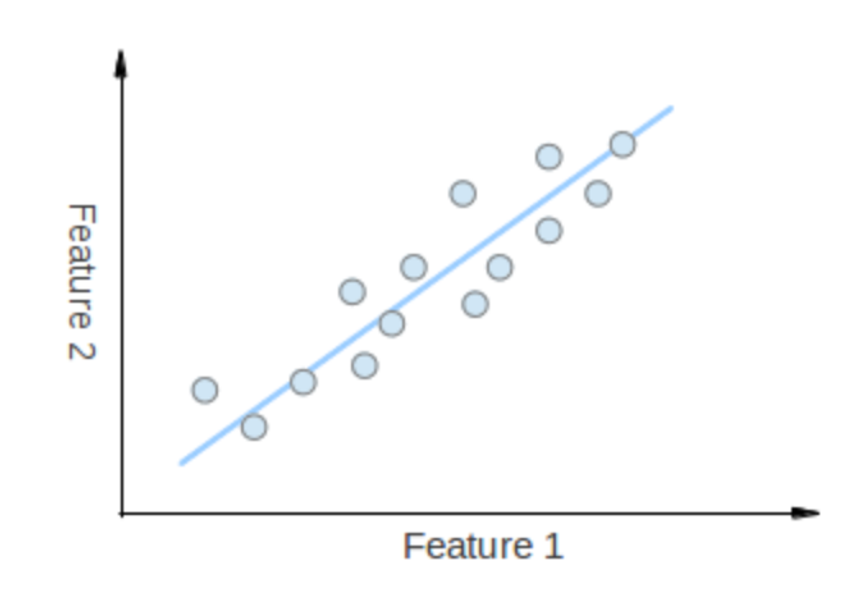
The points vary the most along the blue line, more than they vary along the Feature 1 or Feature 2 axes, so we'll be better off with that dimension only
We want to turn dataset into a small dimension one
Karhunen-Loève transform:
Algorithm
Compute empirical mean for every dimension
, with vector of size (p, 1)
Compute deviation from mean
with , of size (n, 1)
Find the covariant matrix of size (p, p)
, where is the conjugate transpose of
If B consists only of real number, it is just the regular transpose.
SVD decomposition of the covariant matrix to find eigenvalues and eigenvectors
With , with being the i-th eigenvalues of C
are eigenvectors, come in pairs with eigenvalues
full script
Python
Copy
from __future__ import print_function
from __future__ import division
import cv2 as cv
import numpy as np
import argparse
from math import atan2, cos, sin, sqrt, pi
def drawAxis(img, p_, q_, colour, scale):
p = list(p_)
q = list(q_)
angle = atan2(p[1] - q[1], p[0] - q[0]) # angle in radians
hypotenuse = sqrt((p[1] - q[1]) * (p[1] - q[1]) + (p[0] - q[0]) * (p[0] - q[0]))
# Here we lengthen the arrow by a factor of scale
q[0] = p[0] - scale * hypotenuse * cos(angle)
q[1] = p[1] - scale * hypotenuse * sin(angle)
cv.line(img, (int(p[0]), int(p[1])), (int(q[0]), int(q[1])), colour, 1, cv.LINE_AA)
# create the arrow hooks
p[0] = q[0] + 9 * cos(angle + pi / 4)
p[1] = q[1] + 9 * sin(angle + pi / 4)
cv.line(img, (int(p[0]), int(p[1])), (int(q[0]), int(q[1])), colour, 1, cv.LINE_AA)
p[0] = q[0] + 9 * cos(angle - pi / 4)
p[1] = q[1] + 9 * sin(angle - pi / 4)
cv.line(img, (int(p[0]), int(p[1])), (int(q[0]), int(q[1])), colour, 1, cv.LINE_AA)
def getOrientation(pts, img):
sz = len(pts)
data_pts = np.empty((sz, 2), dtype=np.float64)
for i in range(data_pts.shape[0]):
data_pts[i,0] = pts[i,0,0]
data_pts[i,1] = pts[i,0,1]
# Perform PCA analysis
mean = np.empty((0))
mean, eigenvectors, eigenvalues = cv.PCACompute2(data_pts, mean)
# Store the center of the object
cntr = (int(mean[0,0]), int(mean[0,1]))
cv.circle(img, cntr, 3, (255, 0, 255), 2)
p1 = (cntr[0] + 0.02 * eigenvectors[0,0] * eigenvalues[0,0], cntr[1] + 0.02 * eigenvectors[0,1] * eigenvalues[0,0])
p2 = (cntr[0] - 0.02 * eigenvectors[1,0] * eigenvalues[1,0], cntr[1] - 0.02 * eigenvectors[1,1] * eigenvalues[1,0])
drawAxis(img, cntr, p1, (0, 255, 0), 1)
drawAxis(img, cntr, p2, (255, 255, 0), 5)
angle = atan2(eigenvectors[0,1], eigenvectors[0,0]) # orientation in radians
return angle
parser = argparse.ArgumentParser(description='Code for Introduction to Principal Component Analysis (PCA) tutorial.\
This program demonstrates how to use OpenCV PCA to extract the orientation of an object.')
parser.add_argument('--input', help='Path to input image.', default='pca_test1.jpg')
args = parser.parse_args()
src = cv.imread(cv.samples.findFile(args.input))
# Check if image is loaded successfully
if src is None:
print('Could not open or find the image: ', args.input)
exit(0)
cv.imshow('src', src)
# Convert image to grayscale
gray = cv.cvtColor(src, cv.COLOR_BGR2GRAY)
# Convert image to binary
_, bw = cv.threshold(gray, 50, 255, cv.THRESH_BINARY | cv.THRESH_OTSU)
contours, _ = cv.findContours(bw, cv.RETR_LIST, cv.CHAIN_APPROX_NONE)
for i, c in enumerate(contours):
# Calculate the area of each contour
area = cv.contourArea(c)
# Ignore contours that are too small or too large
if area < 1e2 or 1e5 < area:
continue
# Draw each contour only for visualisation purposes
cv.drawContours(src, contours, i, (0, 0, 255), 2)
# Find the orientation of each shape
getOrientation(c, src)
cv.imshow('output', src)
cv.waitKey()
Kmeans
Basic intuition
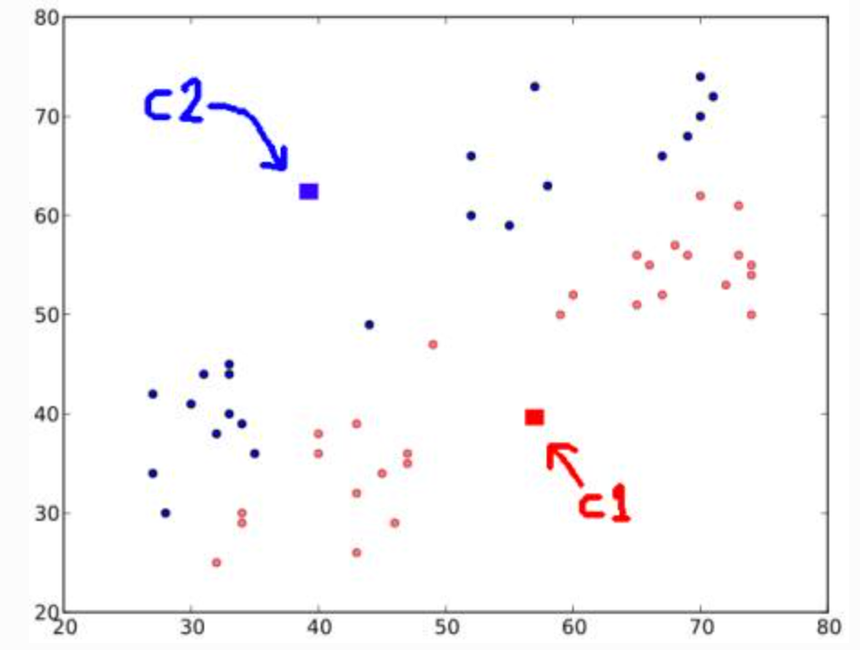
Randomly choose 2 centroids, and
Calculate distance for each from both centroid. Closer to are labeled , o.w labeled
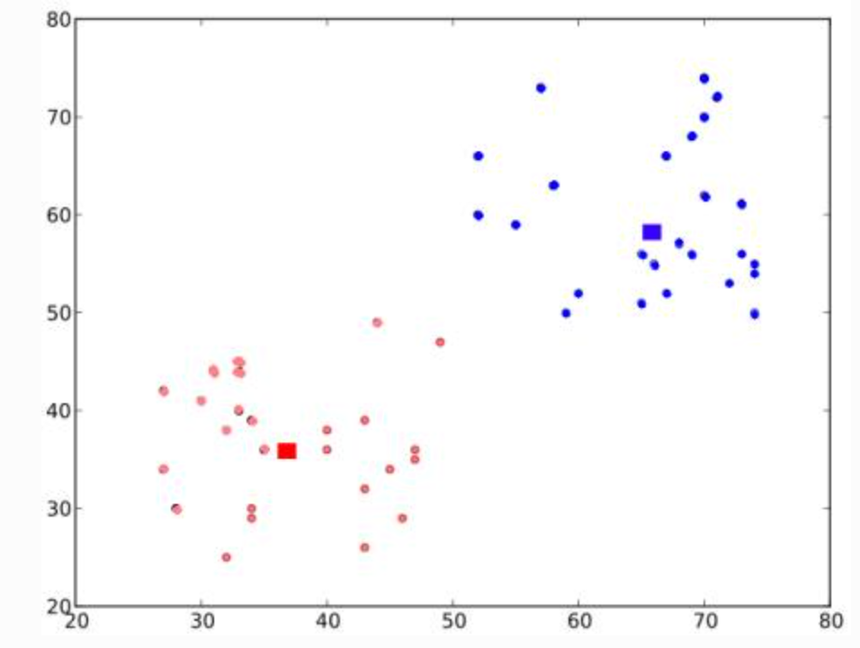
Next, compute the average for blue and red points and update the centroids
Iterate over 2 and 3 until convergence (or stopping criteria on number of iteration, or precision) is reach
The distances between test data and their centroids are minimum
full script
Python
Copy
import numpy as np
import cv2
img = cv2.imread('home.jpg')
Z = img.reshape((-1,3))
# convert to np.float32
Z = np.float32(Z)
# define criteria, number of clusters(K) and apply kmeans()
criteria = (cv2.TERM_CRITERIA_EPS + cv2.TERM_CRITERIA_MAX_ITER, 10, 1.0)
K = 8
ret, label, center = cv2.kmeans(
Z, K, None, criteria, 10, cv2.KMEANS_RANDOM_CENTERS
)
# Now convert back into uint8, and make original image
center = np.uint8(center)
res = center[label.flatten()]
res2 = res.reshape((img.shape))
cv2.imshow('res2',res2)
cv2.waitKey(0)
cv2.destroyAllWindows()